Single-cell RNA sequencing (scRNA-seq) enables high-resolution profiling of individual cells, allowing researchers to identify cell types, track differentiation pathways, and analyze cell fate decisions within organoids.

The human brain’s structural complexity originates from tightly regulated patterning events during early development, where morphogens establish spatial and temporal gradients to define regional identity. However, a systematic understanding of how human neuroepithelial cells respond to these signals has remained limited.
A recent study published in Nature Methods addresses this gap using large-scale single-cell RNA sequencing (scRNA-seq) to profile neural organoid responses to diverse morphogen conditions. By systematically varying timing, concentration, and combinations, the study provides a high-resolution map of how morphogens shape cell fate decisions in vitro.
This work offers a valuable reference framework for optimizing neural organoid models and improving experimental design in developmental and disease research.
To systematically evaluate morphogen responses, the researchers established a high-throughput neural organoid screening workflow in a 96-well format. Human pluripotent stem cell-derived organoids were exposed to signaling factors including SHH, FGF8, retinoic acid, BMP4, BMP7, and WNT pathway modulators under different treatment windows, concentrations, and combinations. After 21 days of differentiation, the organoids were dissociated and analyzed by single-cell RNA sequencing on the 10x Genomics platform, with CITE-seq-based cell hashing used for multiplexing.
Module | Method name | Cell lines used | Main reagents/tools |
Cell culture | hPSC culture | HES3, H9, WTC, H1, WIBJ2 | mTeSR Plus, Matrigel, EDTA |
Organoid induction | Minimal neural induction medium (MNIM) | MNIM, NdiffiVA, heparin | |
Morphogen treatment | Time, concentration, combination screening | Mainly HES3 | CHIR, XAV939, SHH, FGF8, RA, BMP4/7, cyclopamine |
Single-cell sequencing (transcriptome + surface protein) | 10x Genomics + CITE-seq (cell hashing) | Mixed cell lines | 10x Chromium, TotalSeq-A antibodies |
Cell line identity | demuxlet / vireo | Mixed cell lines | demuxlet, vireo, cellsnp-lite |
Data analysis | Seurat + SCENIC | All cell lines | Seurat, pySCENIC, AUCell |
Spatial mapping | VoxHunt | HES3, H9 | VoxHunt, Allen Brain Atlas |
In situ validation | HCR in situ hybridization | WTC, HES3 | HCR probes (Molecular Instruments) |
Microfluidic gradient | MiSTR system | H9 | Microfluidic pump, PDMS chip |
Live imaging | Light-sheet microscopy | HES3 (NKX2-1:GFP) | Viventis LSI light-sheet microscope |
Reproducibility analysis | Multiple cell lines + batches | H1, H9, WTC, WIBJ2 | Parse Biosciences kit |
In total, 100,538 single cells across 97 conditions were analyzed, with extended validation reaching over 200,000 cells.
The study establishes a scalable framework to profile morphogen-driven patterning in neural organoids.
Untreated organoids mainly produced generic neural populations
Morphogen treatments induced region-specific identities, including:
o Retina
Non-CNS lineages (e.g., neural crest, non-neural ectoderm) also emerged under specific conditions
This highlights how controlled signaling inputs can expand organoid diversity beyond default neural states.
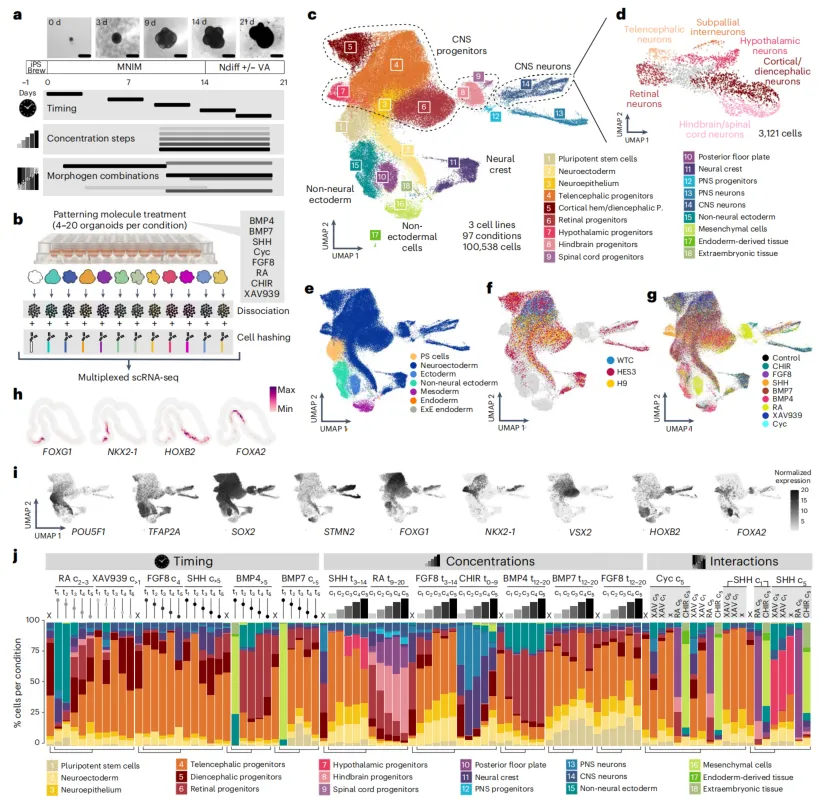
Figure 1.Single-cell transcriptomic atlas of neural organoid patterning
The advantage of single-cell data is the ability to quantitatively assess enrichment or depletion of specific cell types by each treatment. The study found that morphogen effects strongly depend on treatment timing and concentration.
Retinoic acid (RA): An early pulse (days 0–3) mainly induces non-neural ectoderm; late application (days 9–20) at low concentration promotes retinal progenitors, while high concentration shifts fate toward hindbrain and floor plate. HCR in situ hybridization validated these findings.
SHH: An early pulse promotes telencephalon; sustained high concentration gradually induces hypothalamic progenitors, consistent with the in vivo SHH concentration gradient.
FGF8: Promotes telencephalon when applied early, but suppresses it when applied late.
BMP4 vs. BMP7: Both suppress neuronal fate early on, but BMP4 is more effective at inducing retinal fate, whereas BMP7 tends to maintain pluripotency.
WNT signaling: The activator CHIR leads to dorsalization and posteriorization, producing neural crest and non-neural tissues; the inhibitor XAV939 promotes telencephalon.
Beyond cell type composition, the study also examines the regulatory logic underlying these fate decisions. Using SCENIC, the researchers inferred 413 transcription factor regulons and linked them to specific morphogen conditions. This analysis revealed that different signaling environments activate distinct downstream gene regulatory programs.
SHH-associated regulons, including NKX2-1 and TCF7L2, were influenced by both treatment timing and concentration. NKX2-1 activity could be induced across multiple treatment windows but required stronger SHH signaling, whereas TCF7L2 was activated more selectively under later and higher-dose conditions. In contrast, FGF8-associated regulons appeared to respond primarily to concentration rather than timing. BMP-related regulons showed the opposite trend, with activation patterns more strongly influenced by treatment window than by dose.
These results reinforce the idea that morphogens do not simply switch cell states on or off. Instead, they engage regulatory programs in a highly context-dependent way, allowing precise modulation of lineage specification through controlled signaling design.
The study also addresses a practical issue that is highly relevant for organoid research: reproducibility. By expanding the analysis across additional cell lines, induction methods, and biological batches, the authors show that identical morphogen conditions do not always produce identical outcomes across experimental systems.
Cell line-specific differences were substantial. Under WNT inhibition, for example, one cell line generated more neural crest and non-neural ectoderm, while another produced stronger telencephalic outcomes. The choice of neural induction protocol also had a major effect. Dual SMAD inhibition increased neuronal output but introduced greater variability between cell lines, whereas minimal neural induction medium produced a broader range of neural and non-neural identities with better consistency across experiments.
Overall, reproducibility across batches remained reasonably strong, but some protocols were clearly more variable than others. Interestingly, stronger morphogen treatments tended to reduce differences between cell lines, suggesting that robust signaling inputs may partially override intrinsic variability in differentiation potential.
To better approximate in vivo morphogen gradients, the researchers also compared conventional static organoid treatment with a microfluidic gradient model. In the microfluidic system, SHH was delivered across a two-dimensional neural tissue in a continuous gradient rather than as a uniform concentration.
Single-cell integration showed that both systems generated broadly similar cell identities, but their spatial organization differed. The gradient model produced a more continuous transition across regional states, while static organoids often displayed more discrete, jump-like shifts toward strongly ventral identities at particular SHH concentrations. Live imaging further suggested that fate specification in organoids may involve self-organizing feedback, as NKX2-1 expression emerged locally before spreading through the tissue.
This comparison indicates that tissue architecture and mode of signal presentation can meaningfully influence patterning outcomes, even when the same core pathways are involved.
Neural organoid research often involves multi-condition experimental designs, comparative differentiation workflows, and subtle shifts in cell identity that require high-resolution analysis. In these settings, robust single-cell RNA-seq data is essential for accurately profiling cell populations and interpreting developmental trajectories.
Omics Empower provides end-to-end single-cell sequencing support for complex study designs, including library preparation, sequencing, bioinformatics analysis, and publication-oriented data delivery. Our team supports projects across a wide range of tissues and experimental contexts, helping researchers generate reliable datasets for cell type characterization, cell fate analysis, and organoid-based discovery.
Researchers worldwide trust our data: more than 500 peer-reviewed publications have been generated using our single-cell and spatial transcriptomics services, including studies in Nature, Science, and Cell. From library preparation to bioinformatics and publication-ready figures, we deliver end-to-end support to help you advance your next single-cell project.
If you are planning a single-cell sequencing project, these articles may help you evaluate cell sorting strategies, compare platform options, and optimize your experimental workflow from sample preparation to downstream analysis:
Comparing Single-Cell Sequencing Platforms: How to Choose the Right Fit for Your Study
How to Use Flow Cytometry (FACS) Effectively for Single-Cell Sequencing
Is Cell Subtype Annotation Necessary in Single-Cell RNA Sequencing?
Sanchís-Calleja F, Azbukina N, Jain A, et al. Systematic scRNA-seq screens profile neural organoid response to morphogens. Nat Methods. 2026;23:465–478. doi:10.1038/s41592-025-02927-5
Single-cell RNA sequencing (scRNA-seq) enables high-resolution profiling of individual cells, allowing researchers to identify cell types, track differentiation pathways, and analyze cell fate decisions within organoids.
Germany: Arnold-Graffi-Haus / D85 Robert-Rössle-Straße 10 13125 Berlin
United States: (CA) 2 Goddard, Irvine, CA 92618
United States: (IL) 8255 Lemont Rd, #1, Darien, IL 60561
Hong Kong: Unit 615, Building 11W, Hong Kong Science Park, Pak Shek Kok, Hong Kong
Germany: Arnold-Graffi-Haus / D85 Robert-Rössle-Straße 10 13125 Berlin
United States: (CA) 2 Goddard, Irvine, CA 92618
United States: (IL) 8255 Lemont Rd, #1, Darien, IL 60561
Hong Kong: Unit 615, Building 11W, Hong Kong Science Park, Pak Shek Kok, Hong Kong