Xenium Explorer is an interactive visualization and analysis software designed by 10x Genomics specifically for Xenium in situ data. It directly imports output files from the Xenium analysis pipeline—no additional post-processing or format conversion required—making it a fast and convenient tool for exploring spatial transcriptomics datasets.
For researchers working with Xenium datasets, Xenium Explorer provides a practical way to inspect tissue morphology, evaluate cell segmentation, visualize transcript localization, and review clustering results in a spatial context.
In this guide, we walk through the basic workflow, from downloading the software to loading files, navigating the interface, and exporting results. If you are planning a Xenium project and need experimental or analytical support, Omics Empower offers Xenium In Situ service workflows tailored to tissue-based spatial studies.
*All example data and interface images referenced in this article are based on materials from 10x Genomics.
How to Download Xenium Explorer
Visit the official 10x Genomics website and select the appropriate version for your operating system:
Download Xenium Explorer – Official 10x Genomics Support

Required Input Files
To access the full functionality of Xenium Explorer, the required output files should be stored in the same directory before opening the project.
These files typically include:
(1) analysis_summary.html – Analysis summary report
(2) analysis.zarr.zip – Secondary analysis results (e.g., clustering)
(3) cell_feature_matrix.zarr.zip – Expression matrix
(4) cells.zarr.zip – Cell and nucleus segmentation masks for transcript assignment
(5) experiment.xenium – Experimental metadata
(6) gene_panel.json – Gene panel information (panel overview and gene list)
(7) transcripts.zarr.zip – Transcript quality scores and positional information
(8) morphology.ome.tif – DAPI 3D image (nuclear stain)
(9) morphology_focus/morphology_focus_0000.ome.tif – DAPI 2D image
(10) morphology_focus/morphology_focus_0001.ome.tif–Cell boundary image (ATP1A1/E�Cadherin/CD45)
(11) morphology_focus/morphology_focus_0002.ome.tif – Intracellular RNA image (18S RNA)
(12) morphology_focus/morphology_focus_0003.ome.tif–Intracellular protein image (alphaSMA/Vimentin)
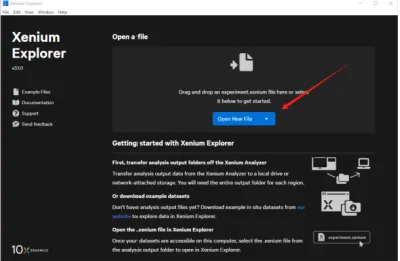
Once the files are in place, open Xenium Explorer, click Open New File, and select the correct folder to load the dataset.
Interactive Interface Features
Xenium Explorer combines tissue image visualization with cell- and transcript-level exploration. Its interface can be understood in three parts: basic navigation tools, image and cell display settings, and transcript visualization options.
Basic Functions
The main toolbar includes several frequently used functions:
Sample Information – View experimental metadata
Image Options – Adjust the field of view
Selections – Use with the lasso tool to select regions of interest
Go to Locations + Save current view – Save a view and later return to it for continued analysis
Mouse Options – Switch between hand (pan) and lasso tools
Export Screenshot – Export the current view as a PNG image
Layer Settings – Three layers (Image, Cells, Transcripts) with adjustable parameters
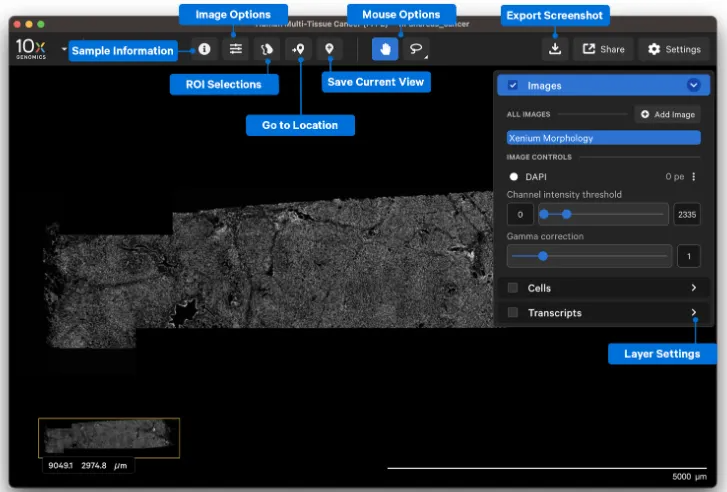
These tools are especially useful when comparing regions, documenting observations, or preparing figures for discussion.
2. Advanced Functions
(1) Images Layer
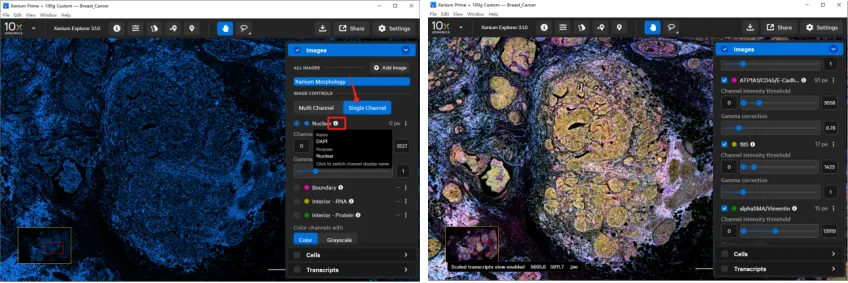
In Xenium Explorer v2.0 and later, the Images layer can load multiple morphology channels from multimodal segmentation datasets. These channels are typically labeled by function, such as:
nuclei
boundary
internal RNA
internal protein
Clicking the information icon next to a channel lets you switch between functional labels and the underlying stain names. Users can also choose to view channels one at a time or combine multiple channels in a multi-channel view.
This makes it easier to compare transcript patterns with tissue structure and staining signals in the same field of view.
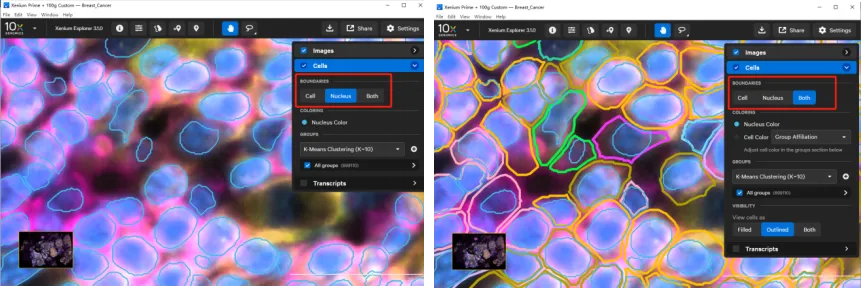
(2) Cells Layer
Under BOUNDARIES, you can show cell outlines, nuclear outlines, or both—a useful feature for assessing cell segmentation quality.
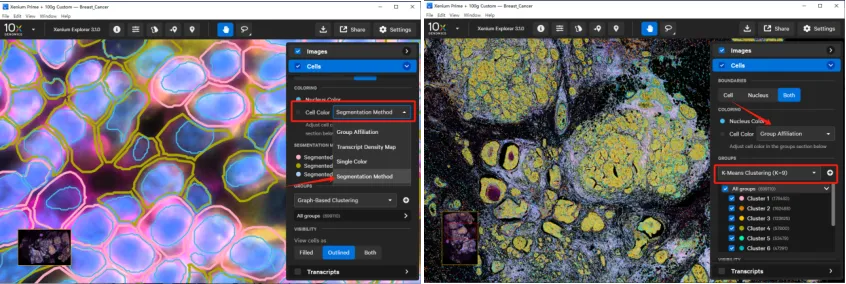
Under COLORING, you can color cells based on clustering results (graph-based, K-means, or custom), transcript density, a single color, or segmentation-related stains. Coloring by cluster helps evaluate whether clustering results align with known tissue types or states.
The GROUPS panel allows you to modify clustering parameters, rename clusters (e.g., change “Cluster1” to a cell type name), and assign colors. You can also upload a custom grouping as a CSV file (with columns for cell_id and custom group) using the “+” button.
(3) Transcripts Layer
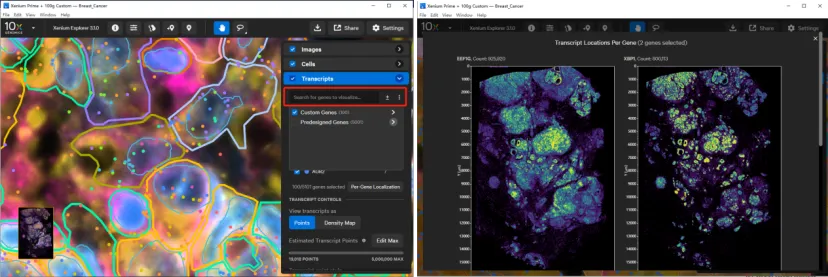
By default, genes are organized by panel origin (e.g., Predesigned Genes and Custom Genes). You can select one or multiple genes to visualize simultaneously across the tissue section. Transcripts can be displayed as dots or as density maps, with dot size and opacity adjustable via the toolbar. Low-quality transcripts (Q-score < 20) are filtered out by default; to show all transcripts, disable Filter out low quality transcripts.
To examine the spatial distribution of a specific gene, search for it in the gene list and then click Per-Gene Localization under the SELECTED GENES panel. This feature can also be used to validate the spatial authenticity of transcript data—for example, by checking genes with known spatially restricted expression patterns.
Result Export and Sharing
1. Exporting ROI and Selected Cell Information
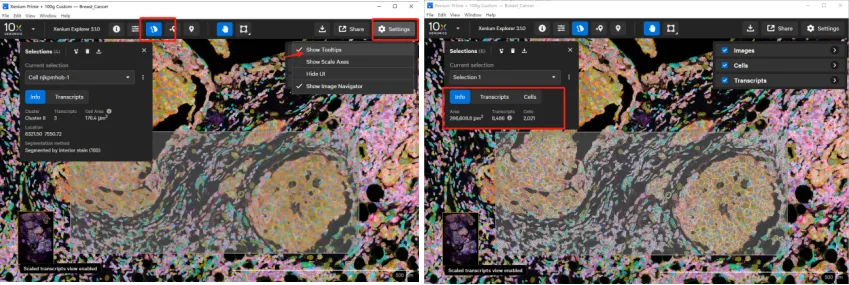
Click the Selections button to select individual cells or regions of interest (ROIs). To select a single cell, ensure Show Tooltips is enabled in Settings. If a selected cell is already within a previously defined ROI, it will not be added as a new entry.
Once a selection is made, the interface displays basic information for the selected area, including ROI boundaries, transcript counts, and cell numbers. For individual cells, detailed metadata and transcript information are provided. These results can be exported as a single CSV package or as individual files.
2. Saving and Sharing View Settings
Click Save Current View to preserve the current image and settings (e.g., zoom coordinates, channel contrast, color selections, opacity, cluster size, scaling thresholds) for later analysis.
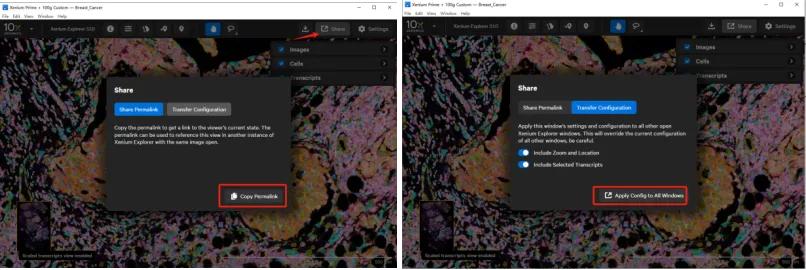
To share settings, click the Share button in the upper right corner. Two options are available:
Share Permalink: Generates a link that can be copied into Go to Location on another computer, transferring the current view settings for collaborative work.
However, this link does not save the following:
selection lists
uploaded custom gene groups
custom transcript colors or symbols
custom cluster names or colors
cell coloring preferences based on grouping
low-quality transcript filter settings
hidden UI states or pop-up windows
Transfer Configurations: Applies the current view settings to all open Xenium Explorer windows on the same computer, overwriting their settings. Like the permalink, this method does not preserve uploaded gene groupings, custom colors, or custom names.
Need Support for Your Xenium Project?
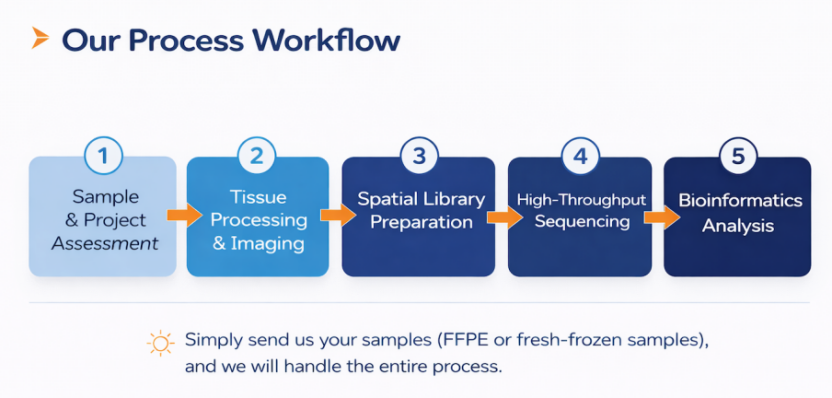
At Omics Empower, we support spatial biology projects with end-to-end workflows, including Xenium In Situ, Visium, and Stereo-seq services. Our team works with researchers across oncology, neuroscience, immunology, and developmental biology to support study design, sample handling, library preparation, sequencing, and downstream analysis.
Why work with us for Xenium projects?
Extensive project experience: Over 20,000 processed samples across neuroscience, oncology, immunology, developmental biology, plant sciences, and agriculture.
Proven research impact: More than 500 publications in single cell sequencing and spatial transcriptomics by our partners in high-impact journals such as Nature, Cell, and the European Heart Journal.
Comprehensive Xenium expertise: Optimized workflows across multiple Xenium platforms, ensuring reliable performance for both human/mouse and non-human projects.
Global laboratory network: With facilities in Europe, Asia, and the United States, we offer consistent service standards, local project support, and faster turnaround times.
Related Articles for Xenium Project Planning
If you are planning a Xenium study, these related articles may help you further evaluate probe design, compare spatial platforms, and choose the most suitable workflow for your research goals: