Xenium is used for targeted spatial in situ analysis in intact tissue sections. It helps researchers study where cells are located, how they interact, and how tissue context shapes biological function.
Single-cell sequencing has fundamentally changed how researchers study cellular heterogeneity. But identifying cell types and states is often only the first step. In many biological questions, researchers also need to understand where cells are located, how they are organized within tissue, and how local interactions shape function or disease.
This is one reason spatially resolved technologies have become increasingly important. Among them, Xenium In Situ from 10x Genomics is gaining strong attention for targeted, high-resolution spatial profiling within intact tissue sections.
In this article, we review three published studies that show how Xenium has been applied in real research settings across cancer, neuroinflammation, and developmental biology. Together, these examples illustrate how spatial in situ analysis can strengthen biological interpretation beyond what dissociated or bulk data can provide.
If you are planning a Xenium project and need experimental or analytical support, Omics Empower offers Xenium In Situ service workflows tailored to tissue-based spatial studies.
Xenium is designed for targeted spatial in situ detection. Using multiplexed fluorescence imaging, it enables the detection of hundreds to thousands of genes directly in tissue sections. Morphological staining of nuclei, cytoplasm, and membranes also supports cell segmentation, making it possible to map gene expression back to individual cells in native tissue architecture.
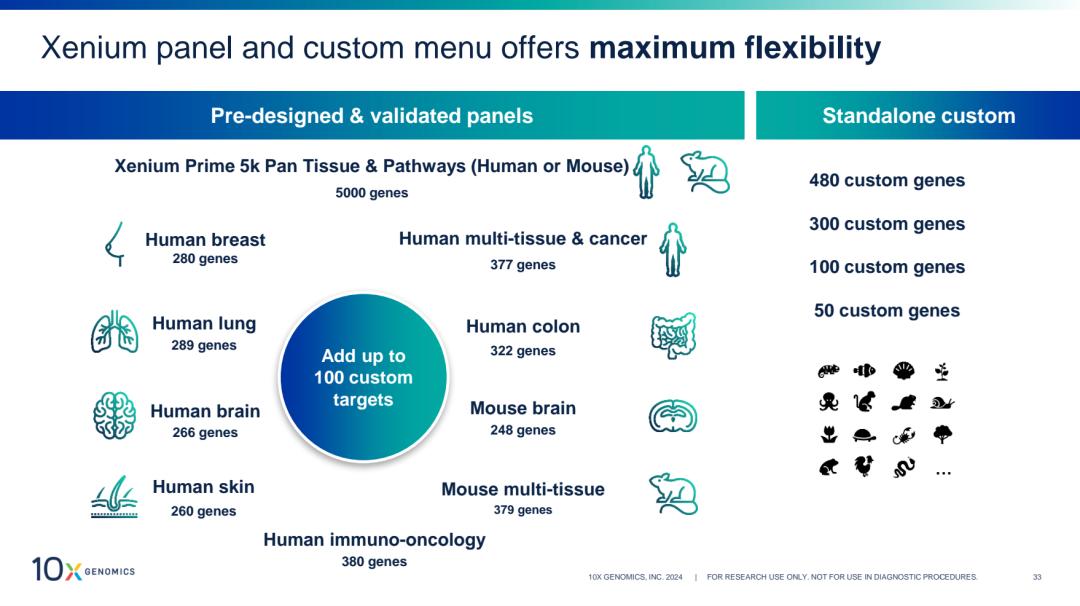
Figure source: 10x Genomics
Several features make Xenium particularly useful in application-driven studies:
Flexible panel options: Xenium 5K Pan-Tissue (5,000 genes) for discovery; tissue-specific panels (200–400 genes) for focused studies.
Custom gene add-on: Up to 100 custom genes per panel—enabling detection of functional markers, novel cell type signatures, mutant sequences, or specific TCR clonotypes.
Subcellular localization: Precise mapping of transcripts enables ligand-receptor interaction analysis, cellular niche characterization, and rare cell identification.
For many studies, Xenium is especially valuable when the goal is not just to identify cell populations, but to determine where they appear, how they interact, and how tissue context influences biological behavior.
Paper: Tumour evolution and microenvironment interactions in 2D and 3D space
Journal: Nature (2024)
Xenium application: Human Pan-Tissue 379-gene + 100-gene custom add-on

Tumor subclones are a major driver of intratumoral heterogeneity and are often associated with immune escape, metastasis, and treatment resistance. In this study, researchers analyzed 78 patient samples across six tumor types and generated Visium spatial transcriptomics data from 131 tissue sections.
They first used CNV-based analysis to identify distinct tumor subclones in different spatial regions. Xenium was then introduced to refine the analysis at higher spatial resolution. By incorporating custom add-on targets for mutated sequences, the authors were able to visualize transcript distributions associated with different subclonal populations directly within tissue.
The study showed that tumor subclones were not evenly distributed. Instead, they occupied distinct spatial niches, supporting the idea that tumor evolution is tightly linked to microenvironmental context.
This example highlights an important application of Xenium in oncology: resolving spatial heterogeneity that may be difficult to interpret using dissociated data alone. It also demonstrates how Xenium can complement broader discovery platforms by validating and refining biologically meaningful spatial patterns.
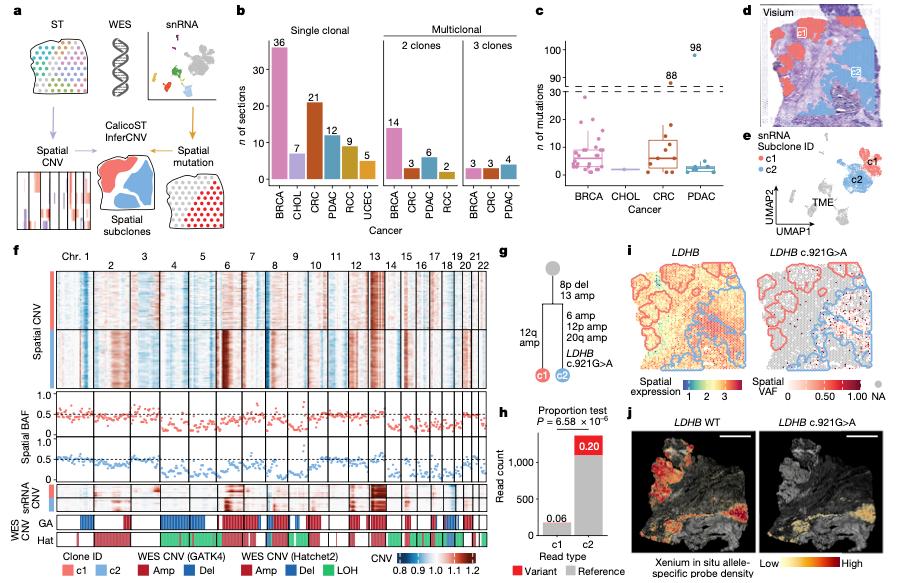
Figure 1. Xenium in situ mapping reveals spatial segregation of tumor subclones.
Paper: Cellular architecture of evolving neuroinflammatory lesions and multiple sclerosis pathology
Journal: Cell (2024)
Xenium application: Human Brain 266-gene panel
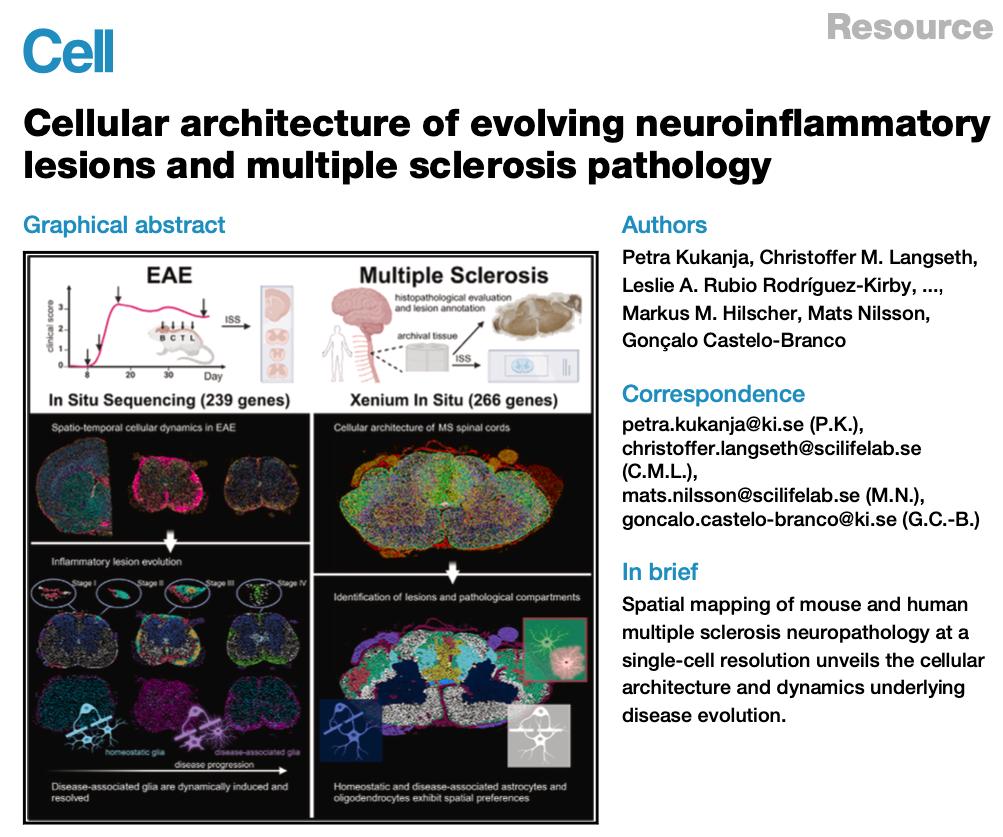
Multiple sclerosis is a chronic neurological disease characterized by multifocal lesions and complex inflammatory pathology. While single-cell analysis has improved our understanding of the cell types involved, it has been harder to capture how these cells are spatially organized during lesion formation and progression.
In this study, the authors used Xenium to investigate both a mouse experimental autoimmune encephalomyelitis model and human multiple sclerosis tissue. They found that immune cells such as T cells and monocytes accumulated in restricted regions during early disease and then spread spatially as pathology progressed.
The analysis also defined multiple lesion-associated regions with different cellular compositions. Monocytes and dendritic cells were enriched in lesion cores, while disease-associated glial cells were more prominent in surrounding areas. By integrating spatial localization with interaction analysis, the study suggested that these glial populations may perform different regulatory functions depending on lesion stage and microenvironment.
This work illustrates how Xenium can support spatial pathology research in neuroinflammatory disease, especially when the aim is to understand lesion structure, immune infiltration, and cell–cell interactions within tissue context.
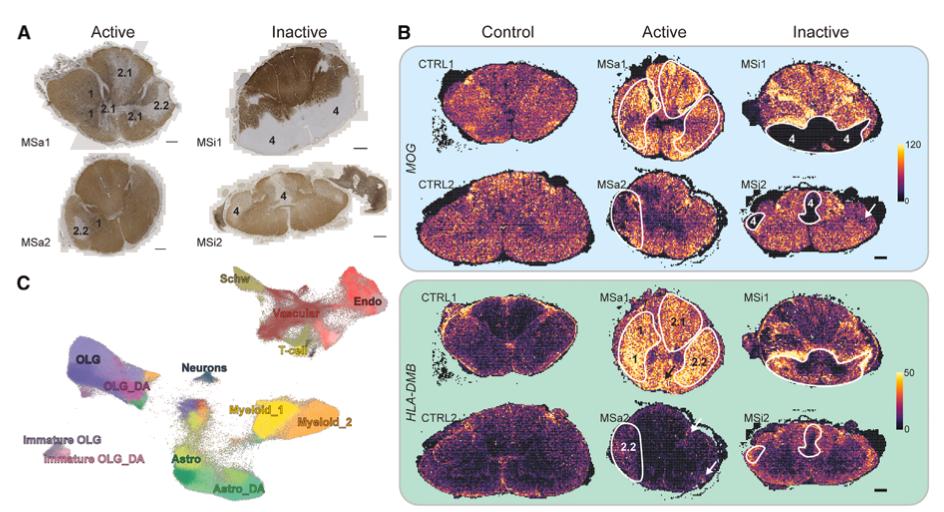
Figure 2. Spatial distribution of disease-associated (DA) glial cells in MS lesions.
Paper: Early human fetal lung atlas reveals the temporal dynamics of epithelial cell plasticity
Journal: Nature Communications (2024)
Xenium application: Human Lung 289-gene + 50-gene custom add-on

Developmental biology often requires both temporal and spatial resolution. In this study, researchers analyzed approximately 150,000 single cells from healthy human fetal lung tissue collected between 10 and 19 weeks of gestation.
Their analysis identified CFTR-positive progenitor populations and reconstructed developmental trajectories toward multiple epithelial cell types. Xenium was then used to place these cell populations back into tissue context, allowing the authors to examine where these progenitors and differentiated descendants appeared over time.
This spatial analysis helped reveal how signaling pathways may regulate the emergence of specialized epithelial populations, including ciliated cells and pulmonary neuroendocrine cells, during lung development.
For developmental studies, this is a strong example of how Xenium can add critical biological context to single-cell data. Rather than showing only what cell states exist, it helps reveal when and where they arise within native tissue structure.
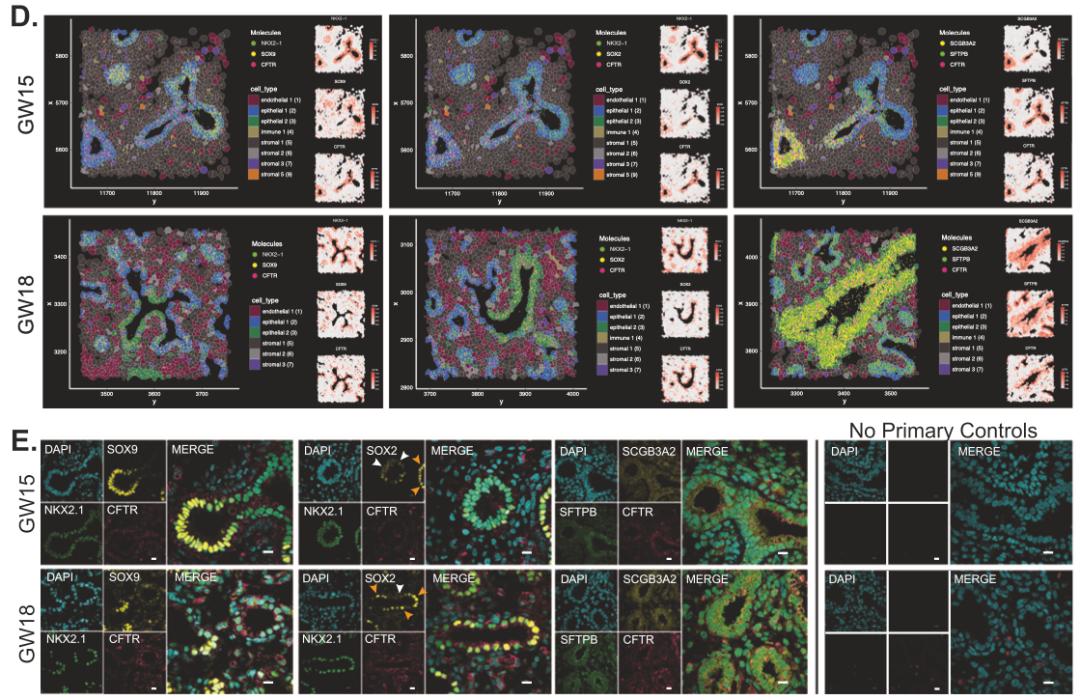
Figure 3. Spatiotemporal dynamics of CFTR+ progenitor cells during human fetal lung development.
Although these three studies focus on different biological systems, they point to a shared theme: Xenium is most powerful when spatial context is central to the research question.
Across oncology, neuroinflammation, and developmental biology, researchers are using Xenium to:
map spatially restricted cell populations
study local tissue organization
investigate cellular neighborhoods and microenvironments
validate findings from single-cell sequencing
refine biological interpretation within intact tissue architecture
This is why Xenium is increasingly appearing in high-impact studies. It is not simply being used as a visualization tool, but as a practical platform for answering biologically meaningful questions that depend on spatial resolution.
At Omics Empower, we support spatial biology projects with end-to-end workflows, including Xenium In Situ, Visium, and Stereo-seq services. Our team works with researchers across oncology, neuroscience, immunology, and developmental biology to support study design, sample handling, library preparation, sequencing, and downstream analysis.

Extensive project experience: Over 20,000 processed samples across neuroscience, oncology, immunology, developmental biology, plant sciences, and agriculture.
Proven research impact: More than 500 publications in single cell sequencing and spatial transcriptomics by our partners in high-impact journals such as Nature, Cell, and the European Heart Journal.
Comprehensive Xenium expertise: Optimized workflows across multiple Xenium platforms, ensuring reliable performance for both human/mouse and non-human projects.
Global laboratory network: With facilities in Europe, Asia, and the United States, we offer consistent service standards, local project support, and faster turnaround times.
If you are planning a Xenium study, these related articles may help you further evaluate probe design, compare spatial platforms, and choose the most suitable workflow for your research goals:
1. Mo CK, Liu J, Chen S, et al. Tumour evolution and microenvironment interactions in 2D and 3D space. Nature. 2024. doi:10.1038/s41586-024-08087-4.
2. Kukanja P, Langseth CM, Rubio Rodríguez-Kirby LA, et al. Cellular architecture of evolving neuroinflammatory lesions and multiple sclerosis pathology. Cell. 2024. doi:10.1016/j.cell.2024.02.030.
3. Quach H, Farrel S, Wu MJM, et al. Early human fetal lung atlas reveals the temporal dynamics of epithelial cell plasticity. Nat Commun. 2024. doi:10.1038/s41467-024-50281-5.
Xenium is used for targeted spatial in situ analysis in intact tissue sections. It helps researchers study where cells are located, how they interact, and how tissue context shapes biological function.
Germany: Arnold-Graffi-Haus / D85 Robert-Rössle-Straße 10 13125 Berlin
United States: (CA) 2 Goddard, Irvine, CA 92618
United States: (IL) 8255 Lemont Rd, #1, Darien, IL 60561
Hong Kong: Room 618, Building 6, Hong Kong Science Park, Pak Shek Kok, Hong Kong
Germany: Arnold-Graffi-Haus / D85 Robert-Rössle-Straße 10 13125 Berlin
United States: (CA) 2 Goddard, Irvine, CA 92618
United States: (IL) 8255 Lemont Rd, #1, Darien, IL 60561
Hong Kong: Room 618, Building 6, Hong Kong Science Park, Pak Shek Kok, Hong Kong