Single-cell sequencing has transformed how researchers define cell types and states. Yet gene expression alone does not explain how those states are established. Many biological processes—development, immune activation, tumor progression, and drug resistance—are regulated upstream at the level of chromatin accessibility.
Single-Cell ATAC-Seq (Assay for Transposase-Accessible Chromatin using sequencing) enables genome-wide profiling of chromatin accessibility at single-cell resolution. By identifying accessible regulatory regions in individual cells, researchers can uncover the regulatory architecture underlying cellular identity and heterogeneity.
In late 2018, 10x Genomics announced broad availability of its Chromium Single Cell ATAC solution, accelerating adoption of a standardized, high-throughput scATAC-seq workflow.
What is scATAC-seq—and what does it add beyond bulk ATAC-seq?
Bulk ATAC-seq averages signals across all cells in a sample. In heterogeneous tissues, that can blur:
scATAC-seq assigns accessibility profiles to individual cells/nuclei, enabling:
Tumor heterogeneity mapping (malignant programs vs immune/stromal states)
Gene regulatory program inference (motifs, co-accessibility, integration)
Lineage and trajectory questions (development, differentiation, activation)
Biomarker and regulatory element discovery (state-specific enhancers/promoters)
How 10x single cell ATAC-seq works (Chromium GEM workflow)
The 10x Genomics Chromium platform enables high-throughput single-cell ATAC-Seq using a microfluidic partitioning system.
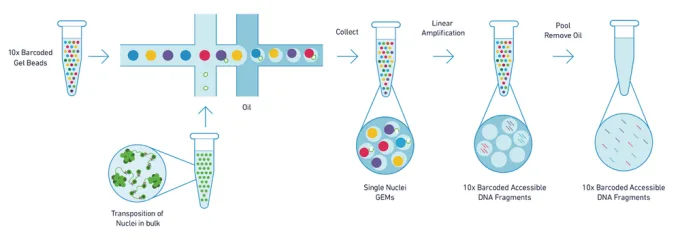
(source: 10x Genomics)
1. Nuclei Preparation and Transposition
Nuclei are isolated and first subjected to transposition (tagmentation), where a transposase preferentially fragments DNA in open chromatin and adds sequencing adapter sequences.
2. GEM Generation and Barcoding
Using microfluidics, individual nuclei are partitioned into GEMs (Gel Beads-in-Emulsion). Each GEM contains a gel bead with a unique barcode. DNA fragments from the same nucleus receive the same barcode, allowing single-cell resolution.
3. Library Construction and Sequencing
Barcoded fragments are amplified and converted into sequencing libraries compatible with sequencing platforms.
4. Computational Processing
Sequencing reads are aligned, filtered, and summarized into accessibility matrices. Downstream analysis typically includes clustering, differential accessibility testing, and transcription factor motif enrichment.
What Can You Learn from Single-Cell ATAC-Seq?
Single-Cell ATAC-Seq helps you understand gene regulation at cell-state resolution by revealing:
Cell-type/state-specific regulatory elements: identify accessible promoters and enhancers that distinguish cell populations.
Putative transcription factor drivers: use motif enrichment in accessible regions to nominate TFs associated with specific states (hypothesis-generating).
Regulation-to-expression links (with scRNA-seq): integrate accessibility with gene expression to prioritize regulatory elements and TF programs most likely shaping observed transcriptional changes.
How Omics Empower supports 10x scATAC-seq projects
Omics Empower provides 10x Chromium–based scATAC-seq as an end-to-end workflow:
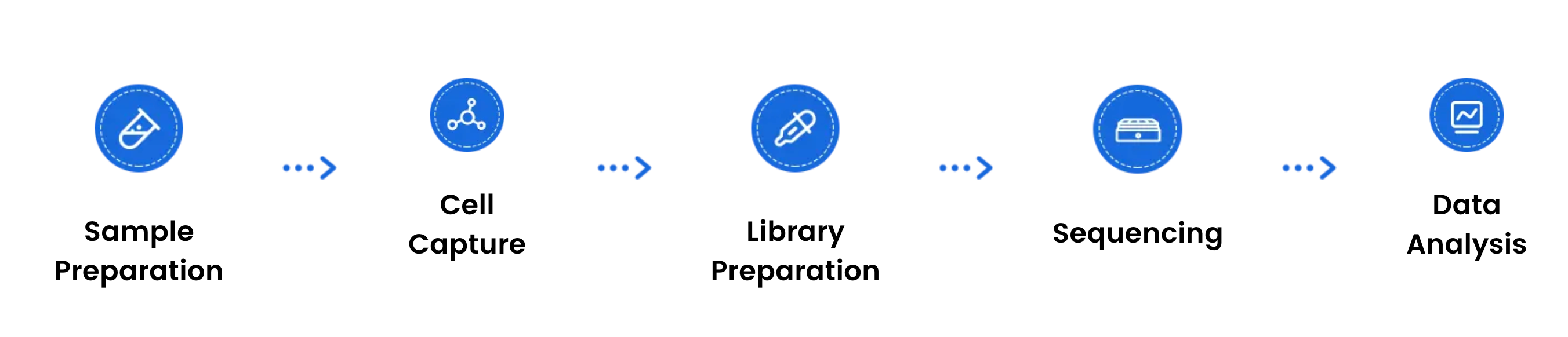
If you’re planning a Single-Cell ATAC-Seq study (or need support with downstream scRNA-seq analysis), working with an experienced service provider can help streamline experimental design, reduce iteration during QC, and speed up time-to-results.
We offer a free consultation to discuss your sample type, study goals, and recommended workflow. You can also review our publication track record to see examples of single-cell sequencing and analysis projects we’ve supported.