Struggling with plant cell type annotation in single-cell sequencing? You’re not alone.
Compared with mammalian systems, plant single-cell analysis often comes with an extra challenge: cell type definitions and marker genes are less standardized, especially for non-model species.
In practice, one of the most common questions researchers ask is:
“How do I confidently define plant cell types in my scRNA-seq dataset?”
A strong starting point is to use curated plant single-cell databases for marker gene lookup, homolog search, and reference-based annotation support.
In this guide, we introduce three widely used plant single-cell resources:
We’ll also show how each database can support your workflow—from marker discovery to cell type annotation, and when it may be helpful to work with an experienced single-cell sequencing service provider.
Why Plant Cell Type Annotation Is Challenging
Plant scRNA-seq has expanded rapidly across roots, leaves, reproductive tissues, and stress-response studies. But annotation remains a bottleneck because:
Marker genes vary across species and tissues
Many non-model plants lack validated markers
Cell states and developmental gradients can blur cluster boundaries
Cross-species annotation often requires homolog-based inference
This is exactly where curated databases become valuable: they help you move from “unknown clusters” to biologically interpretable cell populations.
PlantCellMarker
PlantCellMarker is a user-friendly database that collects plant cell markers from multiple evidence sources, including:
Experimentally validated marker genes
scRNA-seq identified markers
Bulk RNA-seq differential expression markers
Region-specific differentially expressed genes
It covers six model plants:
In addition to marker browsing, PlantCellMarker also provides tools for:
Website: https://www.tobaccodb.org/pcmdb/homePage
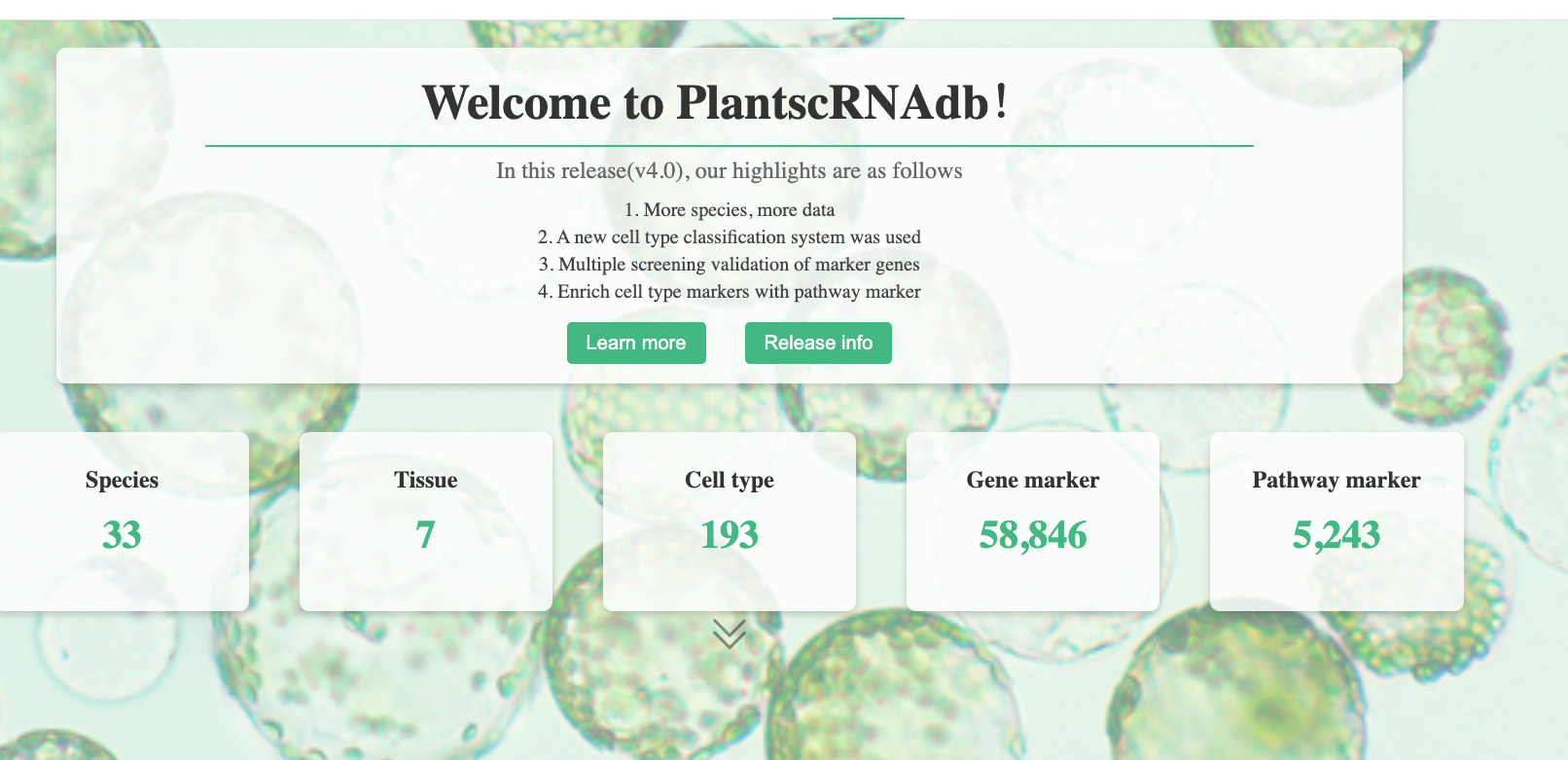
PlantscRNAdb
PlantscRNAdb is a plant single-cell RNA analysis database introduced by a research team from Zhejiang University (published in Molecular Plant in 2021). It includes standardized plant scRNA-seq information across 33 species, such as:
Website: http://ibi.zju.edu.cn/plantscrnadb/#/
Plant Single Cell Transcriptome Hub (PsctH)
Plant Single Cell Transcriptome Hub (PsctH) was reported in Plant Biotechnology Journal in 2021 as an integrated online resource for plant single-cell transcriptome studies.
Unlike marker-only databases, PsctH is designed as a broader end-to-end support platform, with four major components:
Plant single-cell dissociation / protoplast preparation guides
scRNA-seq analysis pipelines
Marker gene resources
Raw plant single-cell sequencing datasets
This makes PsctH especially useful for labs that are new to plant single-cell experiments and need both wet-lab and data-analysis guidance.
Website: http://jinlab.hzau.edu.cn/PsctH/

Need Support for Plant Single-Cell Sequencing
Plant single-cell and spatial transcriptomics projects often involve plant-specific challenges, from sample preparation and tissue handling to data interpretation across complex structures and non-model species.
Since 2018, Omics Empower has been providing single-cell sequencing services and has built extensive experience supporting plant-focused single-cell and spatial transcriptomics research. In 2020, we supported a plant single-cell RNA sequencing study on Arabidopsis thaliana published in Molecular Plant. At the time, this work was among the relatively few plant single-cell transcriptomics studies enabled through collaboration with a research service provider, reflecting early efforts to adapt single-cell technologies to plant systems.
If you are planning a plant single-cell or spatial transcriptomics study—or troubleshooting challenging plant samples—we welcome technical discussions to help define an appropriate workflow and analysis plan.
Reference
1.Chen H, Yin X, Guo L, Yao J, Ding Y, Xu X, et al. PlantscRNAdb: A database for plant single-cell RNA analysis. Mol Plant. 2021;14(6):855-857. doi:10.1016/j.molp.2021.05.002.
2.Xu Z, Wang Q, Zhu X, Wang G, Qin Y, Ding F, et al. Plant Single Cell Transcriptome Hub (PsctH): an integrated online tool to explore the plant single-cell transcriptome landscape. Plant Biotechnol J. 2022;20(1):10-12. doi:10.1111/pbi.13725.