If you have already identified the major cell types in your single-cell RNA-seq dataset, the next question is often more challenging: do you need to go one step further and define cell subtypes?
In our previous guide, Struggling with Cell Type Annotation in scRNA-seq? Here’s Your Essential Guide, we discussed practical strategies for annotating major cell populations in scRNA-seq data. But in many studies, broad cell type labels are only the starting point. To extract deeper biological meaning, researchers often need to further resolve these major populations into biologically relevant subtypes.
This matters because single-cell RNA sequencing is designed to reveal heterogeneity. Major cell type annotation tells you who is present. Cell subtype annotation helps explain how those cells differ functionally, developmentally, or pathologically. In many projects, that second layer is where the real biological story begins.
So, is cell subtype annotation always necessary? Not in every case. But for many publication-oriented or mechanism-driven studies, it is one of the most important steps in single-cell RNA-seq analysis.
Cell subtype annotation refers to the process of dividing a broad cell class into finer, biologically meaningful subpopulations based on transcriptomic features.
For example, after identifying a general immune or epithelial compartment, researchers may further classify cells into more specific subtypes such as CD4 T cells, CD8 T cells, NK cells, activated fibroblasts, secretory epithelial cells, or proliferative progenitor-like states. In reproductive biology, a broader germ-cell lineage might be further resolved into spermatogonia, spermatocytes, and spermatids.
Compared with major cell type annotation, subtype annotation is often more context-dependent. Canonical markers for broad lineages are usually relatively stable, while subtype definitions can vary by tissue, disease setting, developmental stage, and research question. That is why subtype annotation typically requires a combination of marker-gene evidence, clustering results, reference resources, and biological interpretation.
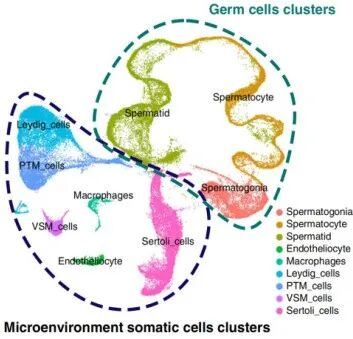
Figure 1. Cell subtype annotation of germ cells and microenvironment somatic cells in single-cell RNA-seq analysis.
In many single-cell RNA-seq studies, major cell type annotation alone is not enough to support strong biological conclusions.
Cell subtype annotation can help researchers:
· Uncover functional heterogeneity within a major lineage
· identify rare or transient populations
· distinguish activation, exhaustion, differentiation, or disease-associated states
· improve downstream analyses such as trajectory inference, cell-cell interaction analysis, and pathway interpretation
· generate more publication-ready biological narratives
In other words, if major cell type annotation provides the basic map of a dataset, subtype annotation often reveals the details that make the map useful.
Not always.
If the goal of a project is only to obtain a broad overview of tissue composition, major cell type annotation may be sufficient in the early stage. However, subtype annotation becomes much more important when:
· the study focuses on cellular heterogeneity
· immune microenvironment analysis is a core objective
· the tissue contains well-defined functional subpopulations
· the project aims to discover novel or disease-associated states
· the data will support a manuscript, grant, or mechanistic conclusion
For many service projects, this is also where analytical depth starts to differentiate a routine result from a truly useful one.
In practice, subtype annotation in scRNA-seq analysis often follows one of two broad strategies: conventional marker-based annotation and non-conventional or cluster-driven annotation.
This is the most common starting point and is especially suitable for cell populations with relatively well-established subtype classifications, such as:
· T and NK cells
· B-cell subsets
· myeloid populations
· trophoblast subtypes
· intestinal epithelial subpopulations
In these cases, the literature often provides canonical markers that can be used to annotate subclusters with reasonable confidence.
The first step is usually to review published marker genes and visualize them using tools such as FeaturePlot, violin plots, dot plots, or Loupe Browser for 10x Genomics datasets. 10x Genomics also provides analysis guidance and annotation-related resources that many researchers use as a practical starting point.
If some clusters cannot be cleanly assigned using canonical markers alone, it is often helpful to consult curated databases and then validate candidate identities back in the dataset. Useful resources include PanglaoDB for cell-type markers, CellMarker 2.0 for curated tissue- and cell-type marker collections, and Cell Taxonomy for broader marker and cell-type reference information. These resources are widely used because they organize marker knowledge across tissues, species, and conditions.
If a cluster remains difficult to interpret, it may not represent a meaningful subtype at all. Researchers should assess low UMI counts, poor-quality cells, doublets, or possible contamination from other lineages before assigning a subtype label. 10x Genomics explicitly notes that mixed or diffuse clusters may require subclustering and evaluation of whether they represent doublets or novel cell types.
If a cluster still cannot be confidently defined, it is better to use labels such as unknown, other epithelial cells, or another lineage-restricted umbrella term rather than overstate the biology.
This marker-first approach is usually the most efficient path when well-accepted subtype definitions already exist.
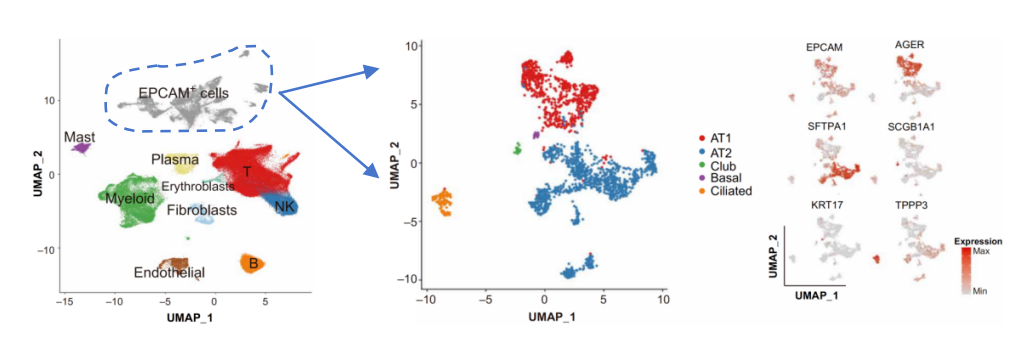
Source: Xing et al. [1].
T and NK cells are a classic example of hierarchical subtype annotation in single-cell RNA-seq analysis.
A common starting point is:
· T cells: express CD3D, CD3E, and CD3G
· NK cells: typically lack coordinatedCD3expression but express markers such as NKG7 or NCAM1
Within the T-cell compartment, researchers often further separate:
· CD8 T cells: express CD8A and/or CD8B
· CD4 T cells: express CD4
· γδ T cells: may express TRDC while lacking typical CD4/CD8 signatures
· NKT cells: may combine T-cell features with NK-related markers such as NKG7 or NCAM1
If a cluster fits neither classic T-cell nor NK-cell patterns, it may be worth testing whether it resembles an ILC-like population, depending on tissue context and marker support.
Researchers should then:
· check sequencing/quality metrics
· review top marker genes
· compare with public references
· apply a cautious label if evidence remains insufficient
The key point is that subtype annotation should move from known biology, to data validation, to conservative interpretation.
Not all cell populations have standard subtype definitions.
In tumor ecosystems, stromal biology, developmental systems, and disease-specific contexts, clusters often represent transcriptional states that do not map neatly onto classical labels. In these situations, a more flexible annotation strategy is needed.
This approach is useful when:
· the literature does not offer stable subtype categories
· the biology is highly context-specific
· the goal is to identify novel cell states
· canonical markers fail to cleanly separate clusters
Researchers often begin by reviewing markers reported in similar tissues, diseases, or model systems.
When published markers do not clearly resolve the clusters, it is acceptable to use the clustering result itself as the working subtype framework.
Common naming strategies include:
Examples include labels such as CCR6+ CD8 T cells, SPP1+ macrophages, or MKI67+ proliferating cells.
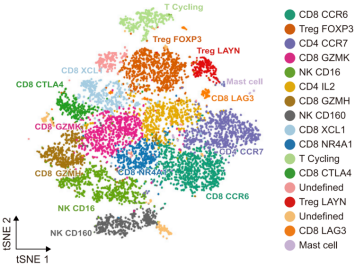

Figure 2. Example of naming cell subtypes based on top marker genes in single-cell RNA-seq analysis. Source: Sun et al. [2].
Some studies retain labels such as C1, C2, C3, especially when the biological identity is still preliminary but the clusters are reproducible and relevant.
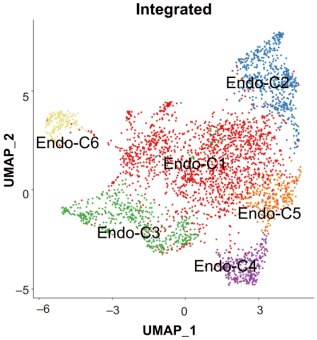
Figure 3. Example of cluster-based subtype naming in single-cell RNA-seq analysis. Source: Xing et al. [1].
If a subtype is defined more by pathway or state signatures than by classical lineage markers, function-based labels may be appropriate, such as interferon-responsive, proliferative, matrix-remodeling, or antigen-presenting populations.
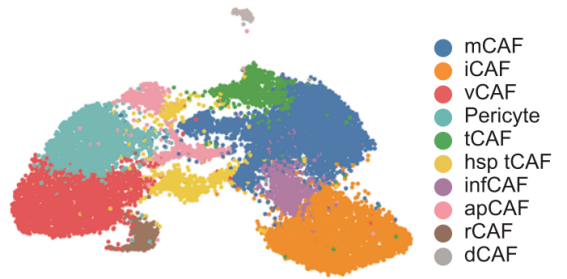
Figure 4. Example of function-based subtype classification in single-cell and spatial proteomics data. Source: Cords et al. [3].
The community best-practices literature on single-cell annotation also emphasizes that cell annotation is the process of labeling groups of cells based on known or sometimes unknown cellular phenotypes, which is exactly why cluster-driven interpretation remains important in real datasets.
If you are planning a single-cell sequencing project and want to generate high-quality data for reliable cell type and cell subtype identification, Omics Empower can support your research with professional single-cell sequencing services.
Researchers worldwide trust our data: more than 500 peer-reviewed publications have been generated using our single-cell and spatial transcriptomics services, including studies in Nature, Science, and Cell. From library preparation to bioinformatics and publication-ready figures, we deliver end-to-end support to help you advance your next single-cell project.
To learn more about cell type annotation, single-cell sequencing, and spatial transcriptomics analysis, explore the following articles from Omics Empower:
· Struggling with Cell Type Annotation in scRNA-seq? Here’s Your Essential Guide
· Plant Single-Cell Sequencing: Cell Type Annotation Databases Guide
· Spatial Transcriptomics Analysis Without Single-Cell Data
Xing X, Li F, Huang Q, et al. Decoding the multicellular ecosystem of lung adenocarcinoma manifested as pulmonary subsolid nodules by single-cell RNA sequencing. Sci Adv. 2021;7:eabd9738. doi:10.1126/sciadv.abd9738.
Sun Y, Wu L, Zhong Y, et al. Single-cell landscape of the ecosystem in early-relapse hepatocellular carcinoma. Cell. 2021;184(2):404-421.e16. doi:10.1016/j.cell.2020.11.041.
Cords L, Tietscher S, Anzeneder T, et al. Cancer-associated fibroblast classification in single-cell and spatial proteomics data. Nat Commun. 2023;14:4294. doi:10.1038/s41467-023-39762-1.
Germany: Arnold-Graffi-Haus / D85 Robert-Rössle-Straße 10 13125 Berlin
United States: (CA) 2 Goddard, Irvine, CA 92618
United States: (IL) 8255 Lemont Rd, #1, Darien, IL 60561
Hong Kong: Room 618, Building 6, Hong Kong Science Park, Pak Shek Kok, Hong Kong
Germany: Arnold-Graffi-Haus / D85 Robert-Rössle-Straße 10 13125 Berlin
United States: (CA) 2 Goddard, Irvine, CA 92618
United States: (IL) 8255 Lemont Rd, #1, Darien, IL 60561
Hong Kong: Room 618, Building 6, Hong Kong Science Park, Pak Shek Kok, Hong Kong